|
Where is the image_tile_size that I can set it to the approximate size of each object. So I need a setting that makes the cells so small so I can see them in the thumbnail. I have also changed the size of the thumbnails to 200 percent and still can not see the cell border but only the size of the thumbnail itself will be large. So how can I make adjustments that allow me to see a single cell directly on each thumbnail selected by the classifier without having to double click on the thumbnail to see the selected cell? Abstract Summary: CellProfiler Analyst allows the exploration and visualization of image-based data, together with the classification of complex. So I need to see the border of the selected cell in the thumbnail so I can select those that are of interest and add them in the positive box. The CellProfiler Analyst 2.0, completely rewritten in Python, builds on these features and adds enhanced supervised machine learning capabilities (Classifier), as well as visualization tools to overview an experiment (Plate Viewer and Image Gallery). Because the cells are so large then I can not see them in the thumbnail selected by classifier but I see a part of the cytoplasm in the selected cell instead of the cell border. Membrane integrity is the feature most often used to detect whether eukaryotic cells cultured in vitro are alive or dead. Now I can understand what the problem is in my case. And it is also important for me to know how to create the property file correctly. I need to first use cell profiler to create a property file and then use that property file in cell profiles analyst to learn and train the program to look for the cells I am interested in. Then I want to train the cell profiler analyst program to look for the elongated cells and quantify them in both treated cells and untreated cells so that I get an idea if the actual number of cells that are elongated is more in the treated cells than in the untreated cells (because I have seen such long cells in untreated cells too, but much less than I have seen in treated cells) then I can conclude that my substance has affected the morphology of the cells and that the appearance of the cells has changed after treatment with the particular substance. You see in the middle of the picture some long and elongated cells and I first want to use cell profiles for the images to be analyzed and segmented as you said and cell profiles can distinguish between the nuclei which are colored in blue and the actin filaments which are colored in red. For applying the machine learning tool to identify a phenotype of interest, watch the tutorial on Classifying Cells with Machine Learning. here I describe my project and also attach a picture of my cells then you can get a better idea of what is important for me to be able to do with those programs. Provides an example of selecting phenotypes using gating. 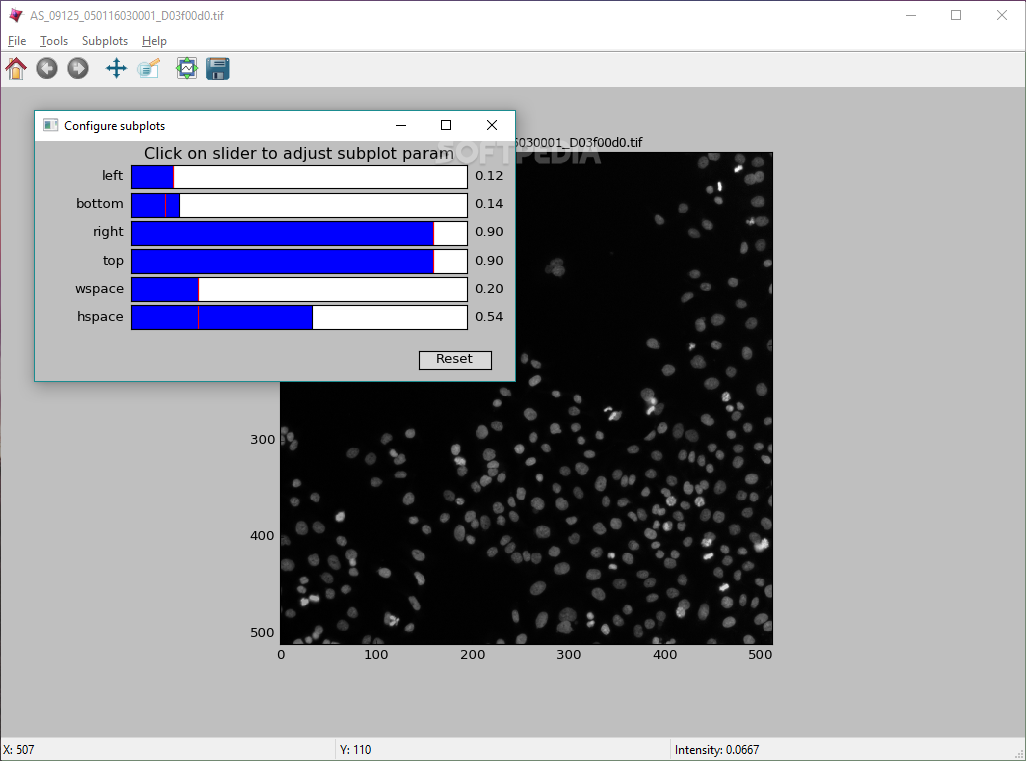 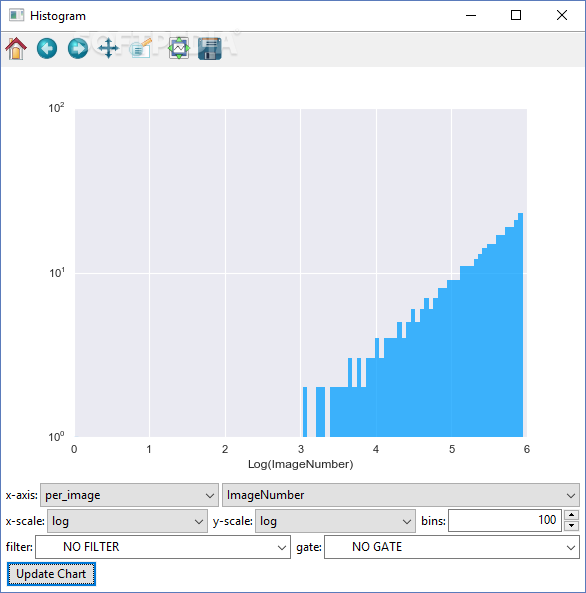
But still, it’s a little hard to know how to use the program for my purpose. I have watched the Translocation tutorial and learned a lot from it.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
